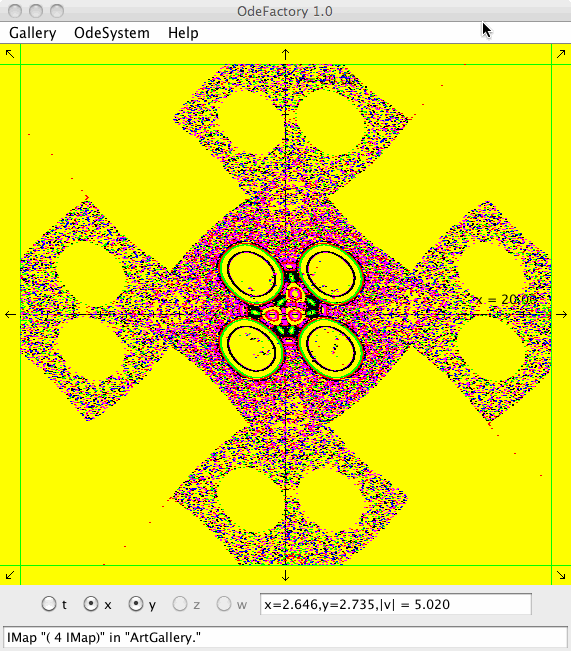
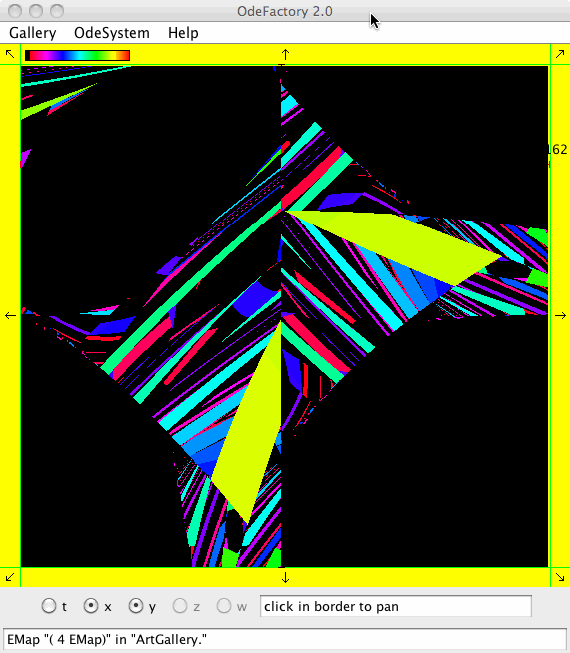
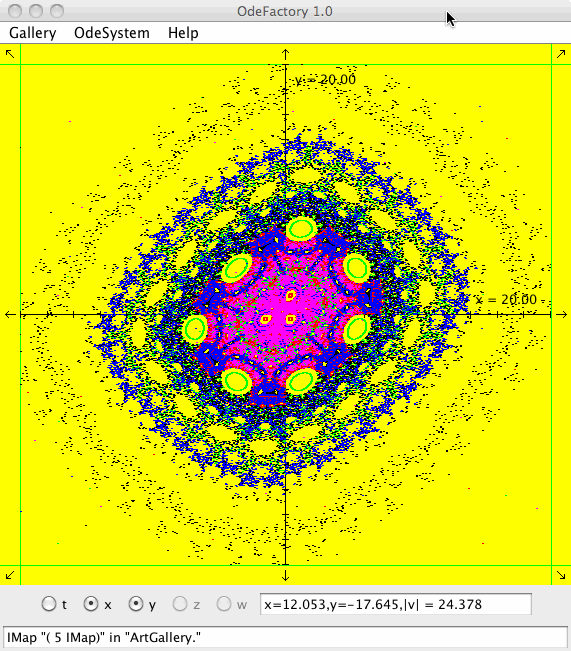
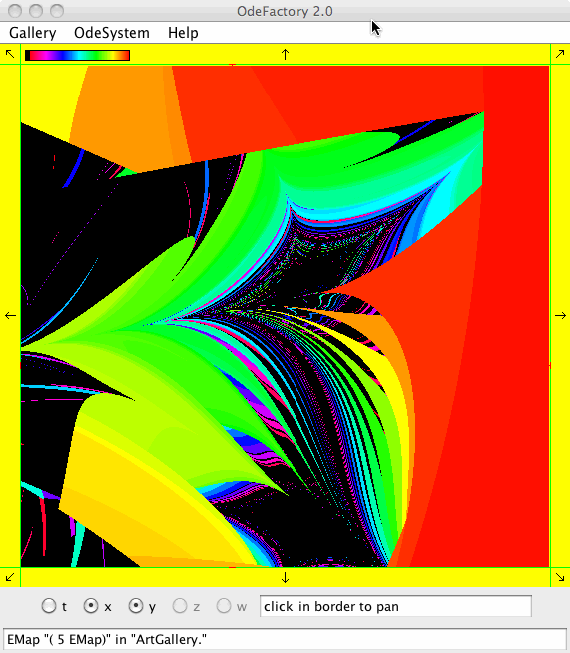




Each vector field defined in OdeFactory can be used to generate graphic Algorithmic art images. The imagers can be saved in jpg files.
A dynamical system defined by a 2D vector field can be viewed as: an Ode, an IMap or a EMap. In the Ode view, you can display: the vector field itself (with the color on or off), trajectories in R2 or R2+, solution curves in 3D, and, components of the solution curves in the (t,x) and (t,y) planes. In the 2D view, you can also zoom in/out and select various parts of images. When a system has parameters, you can generate variations of an image by changing parameter values. For EMap images you can use various color tables and a circular or square prison.
To scroll through 2020 OdeFactory art images click here. Depending on the speed of your internet connection, it may take a while for the images to download from the web. If you scroll too fast there may have a time delay between images.
To view an automatic version of the slide show click here. The automatic slide show displays each of the 2020 images for 5 seconds. It takes about 2.8 hours to complete the slide show.
A hundred older images, from 2013, are shown below. Mouse-over and click on an image for a larger view.
The images are labeled as follows:
view "system name" in "gallery name."
For example, the first image is labeled:
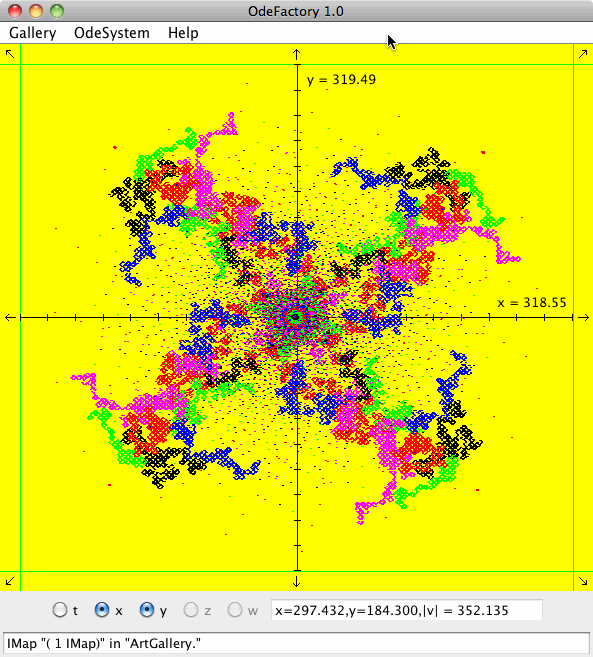
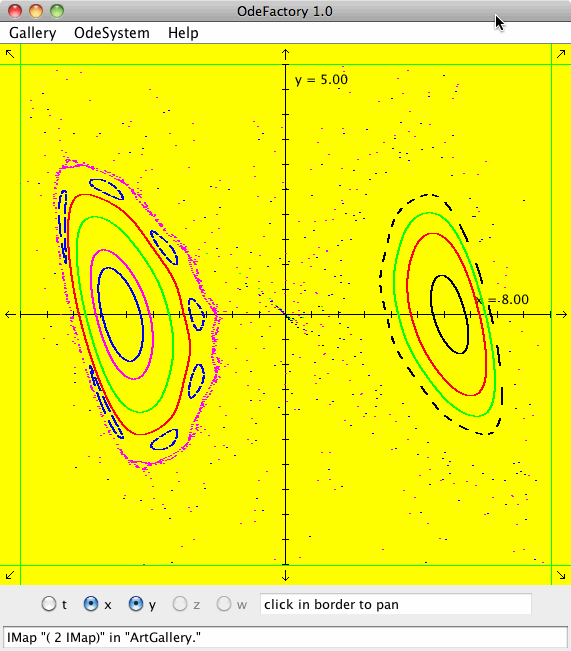
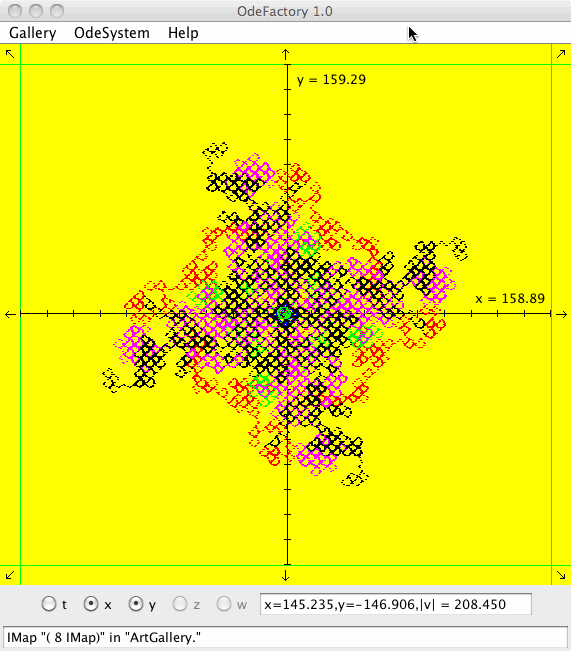
IMap "( 1 IMap)" in "ArtGallery."
meaning you are looking at the IMap view of system "( 1 IMap)" in gallery "ArtGallery." The system is:
x <- y-cos(abs(x)),
y <- a-x.
The older images on this page were originally created between 8/25/12 and 4/21/13. Since then some of the galleries have been updated and the labels on some images have changed. For example, the system name for the first image is now "IMap0011 1" not "( 1 IMap)".
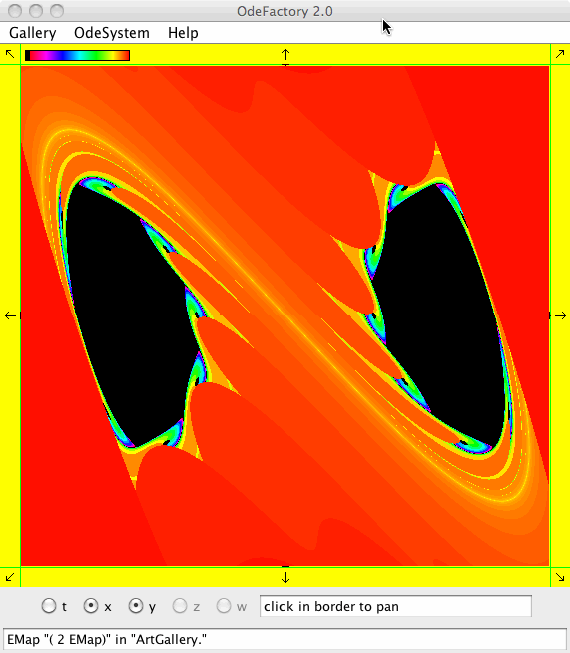
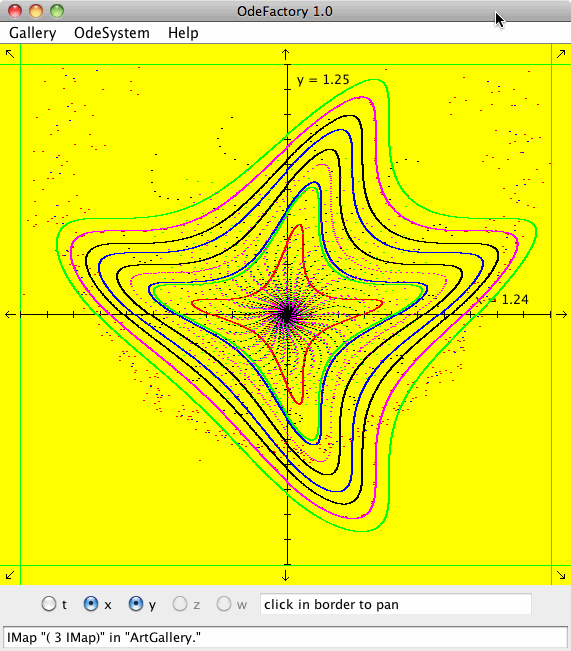
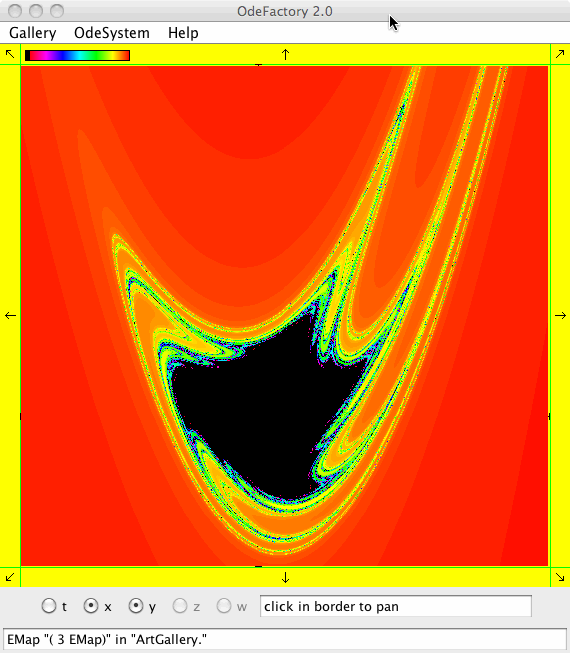
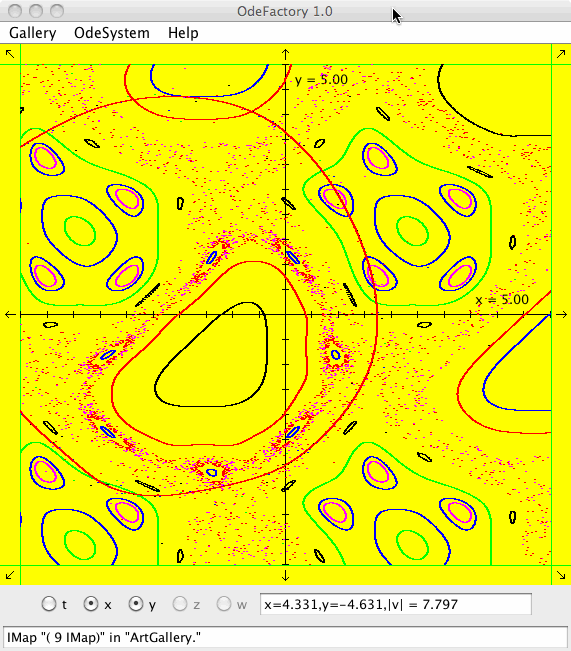
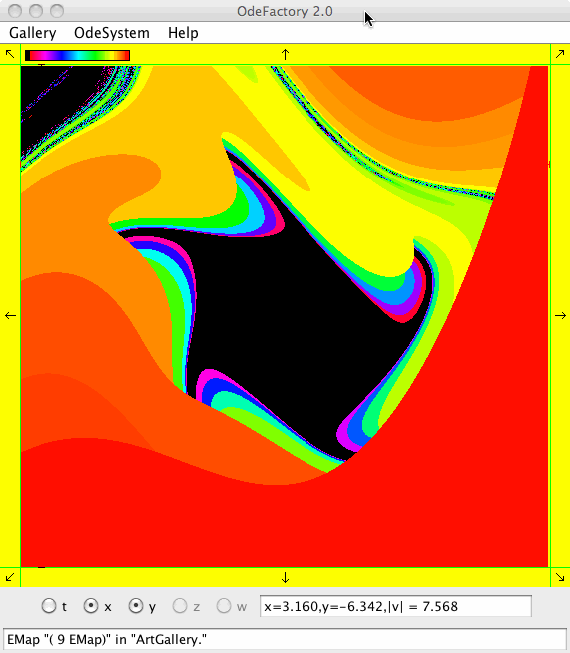
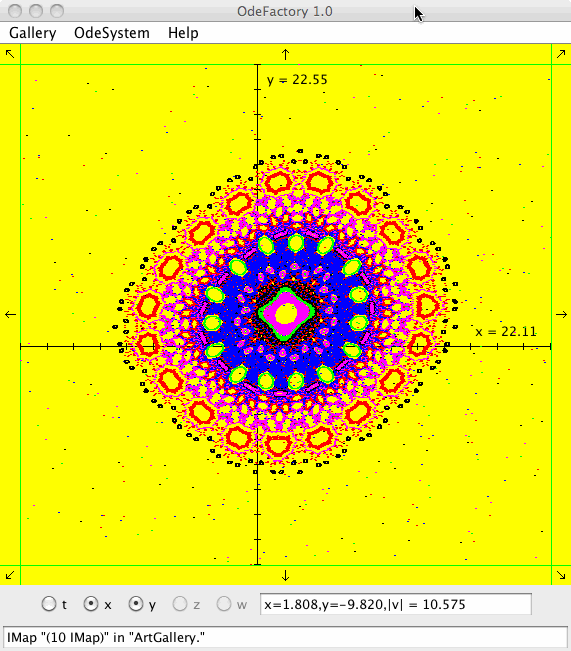
Each of the images is defined by a 2D parameterized vector field. Images labeled "IMap" are views of the iterative map defined by the vector field and images labeled "EMap" are views of the vector field rendered using the escape time algorithm. The floral and/or self-replicating EMap images are fractals and can properly be called “Julia set” images, the more geometric non self-replicating EMap images are not fractals. The one image labeled "Ode" is that of representative trajectories of the ode defined by the vector field.
Variations of the images can be created by adjusting the parameters using OdeFactory. If you want to modify an image, say "( 1 IMap)" in gallery "ArtGallery", download a copy of OdeFactory then double-click on the downloaded OdeFactory.jar file to open OdeFactory. Select item "Open Examples. . ." on the Gallery menu and open gallery "ArtGallery" and then select system "( 1 IMap)" in the gallery. The system opens in the Ode view. Toggle the "IMap" button to get to the IMap view. Zoom in several times and then adjust the parameter "a" to get new variations of the image.
Not all parameter values give interesting images. Generally a system will not be interesting as an "art" image in all three views.
To examine the fractal nature of the EMap images corresponding to fractals, such as system "power flow w/params, p = .55, q = 1" in gallery "EMapExs" use ctrl-move in the viewing area to create a selection square then click the "Update" button in the "Graphics Settings" window to center and zoom in.
You can copy any of the images to your desktop by right-clicking on the image in your browser.
This next set of images are EMap views of systems in various OdeFactory galleries. The system name and gallery name is in the text field at the bottom of each image.
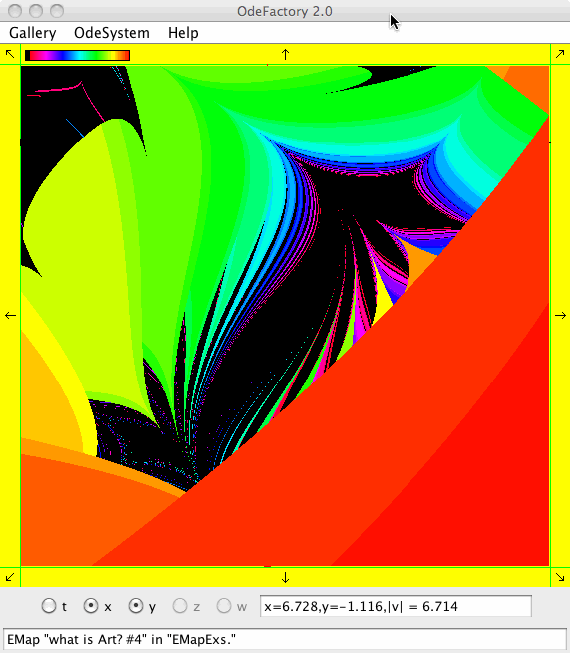
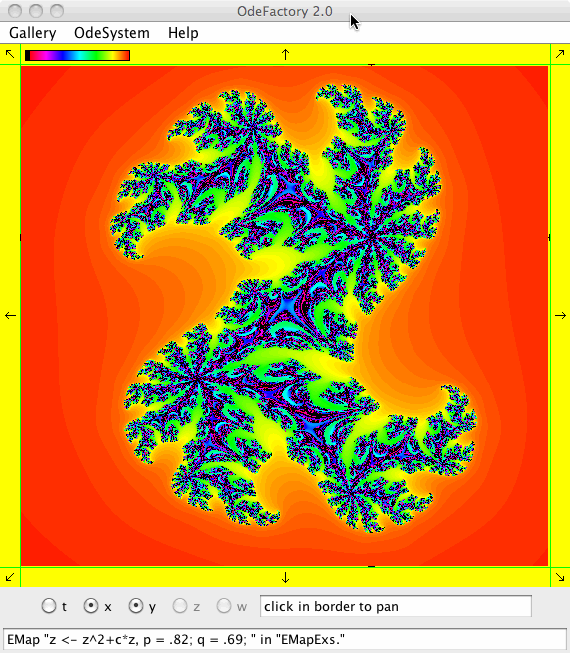
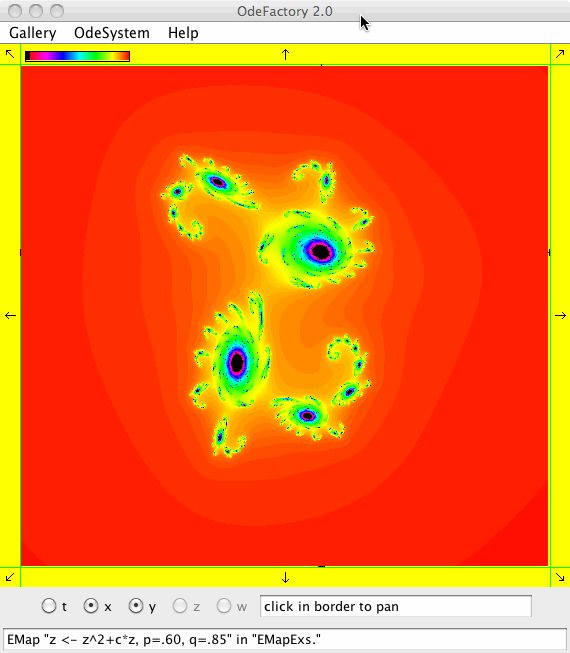
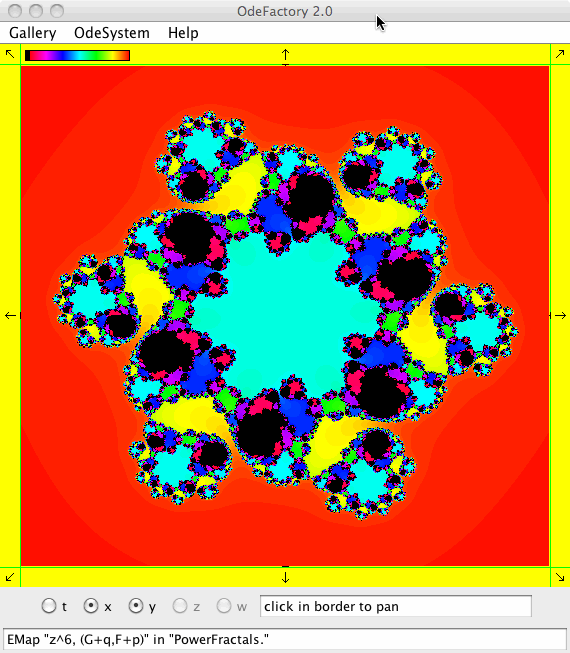
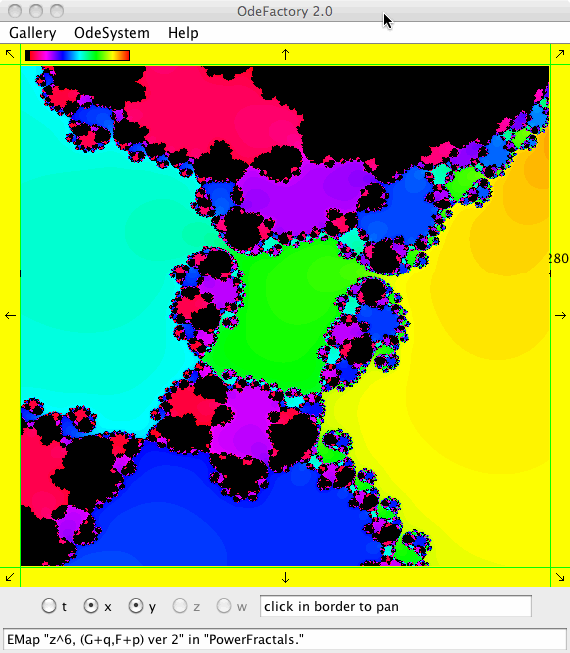
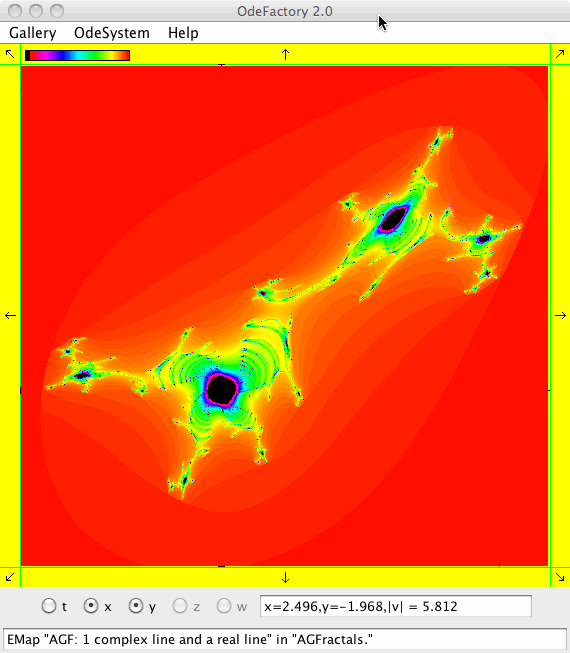

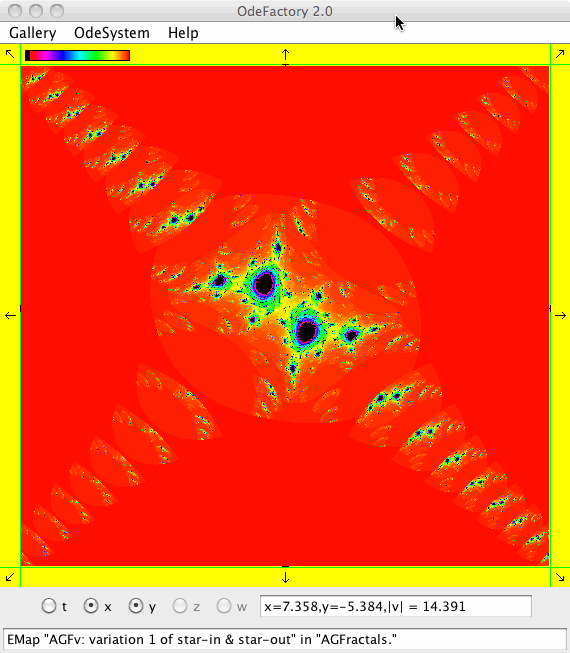
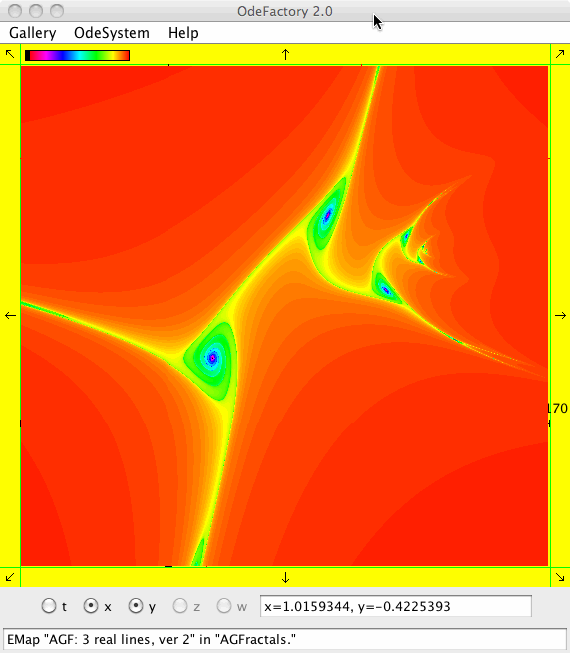
This next set of images are EMap and IMap views of the first 10 systems in OdeFactory gallery HopalongEMaps based on the hopalong map of Barry Martin:
x <- y-sgn(x)*sqrt(abs(b*x-c)),
y <- a-x.

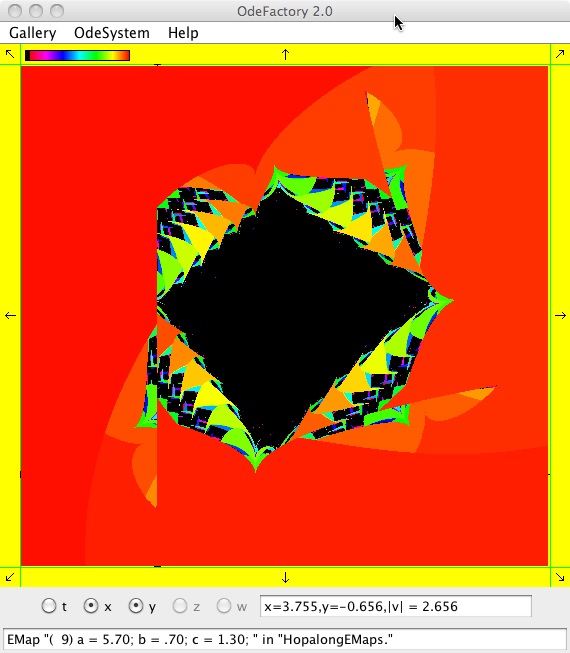
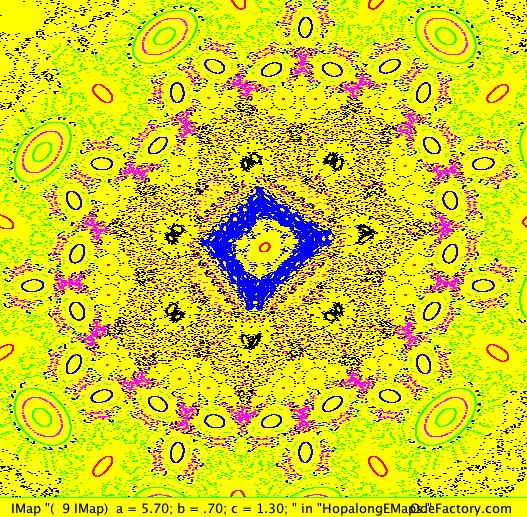
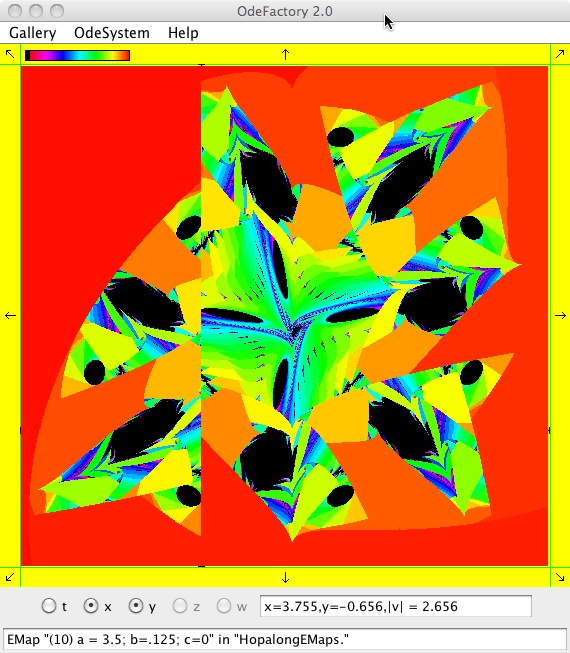

The next 16 images are based on a slight variation of the hopalong map where the second iteration, y<-a-x, is replaced by y<-a-x%d where "%" is the Java remainder or modulus operator which applies to floats as well as to ints.

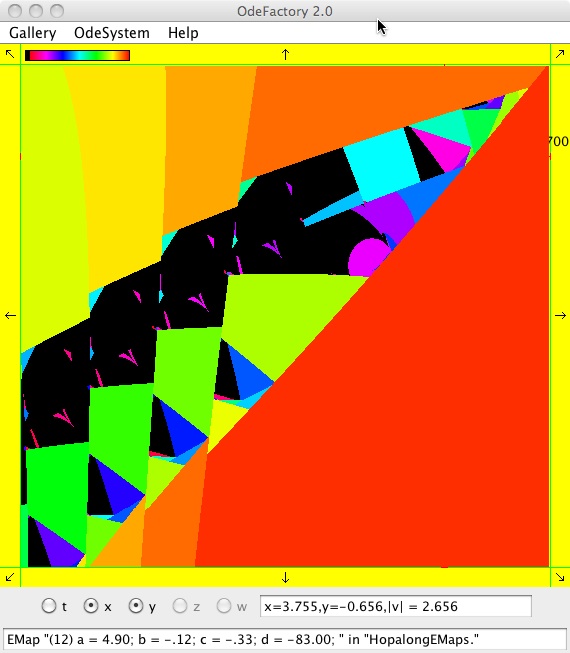
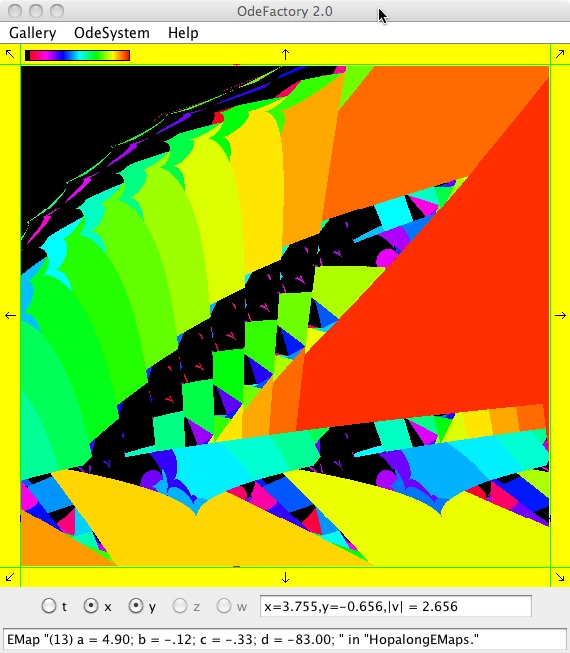
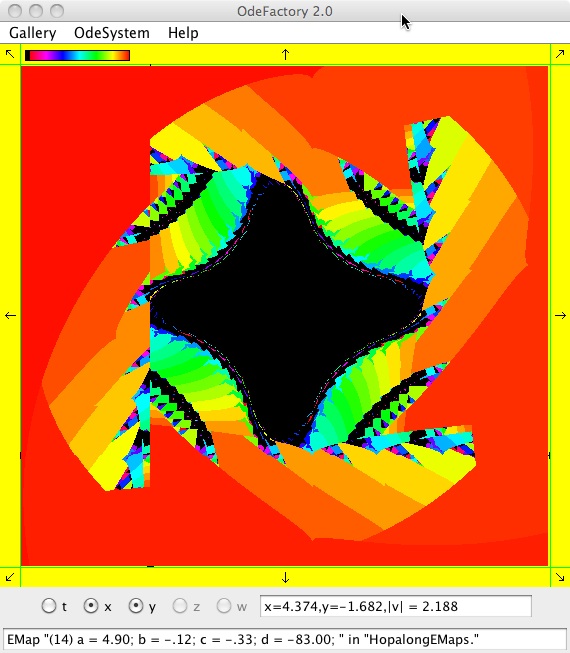
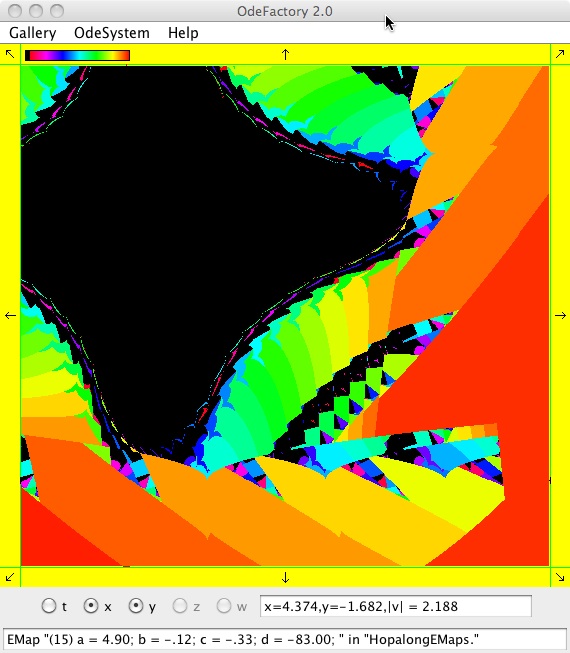

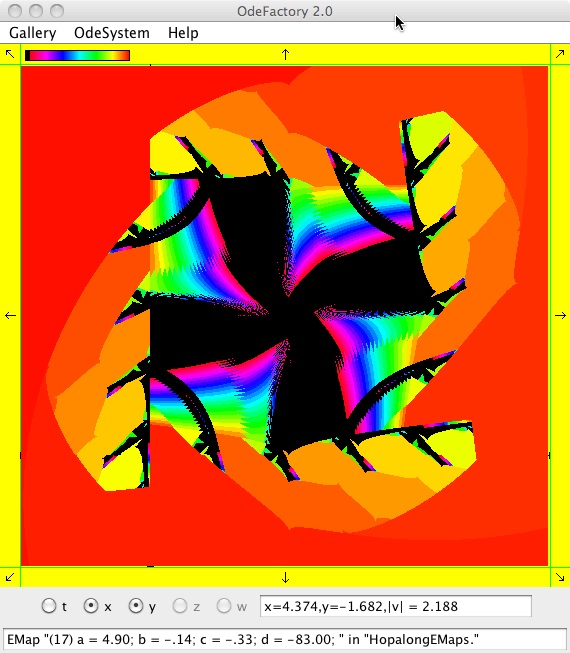
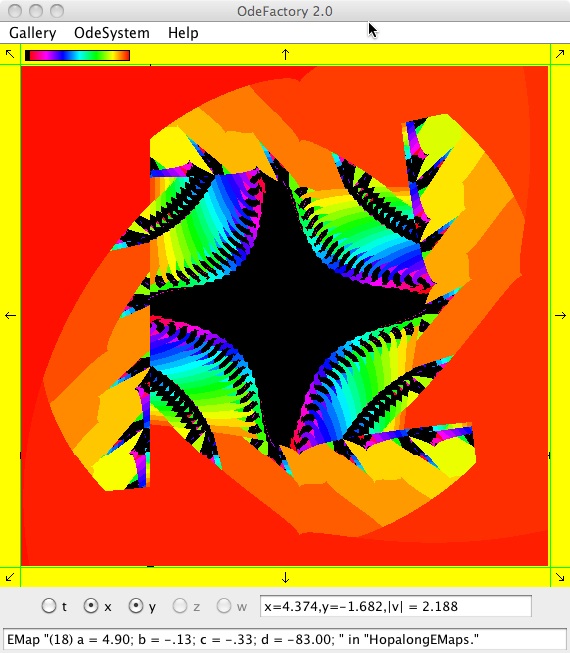
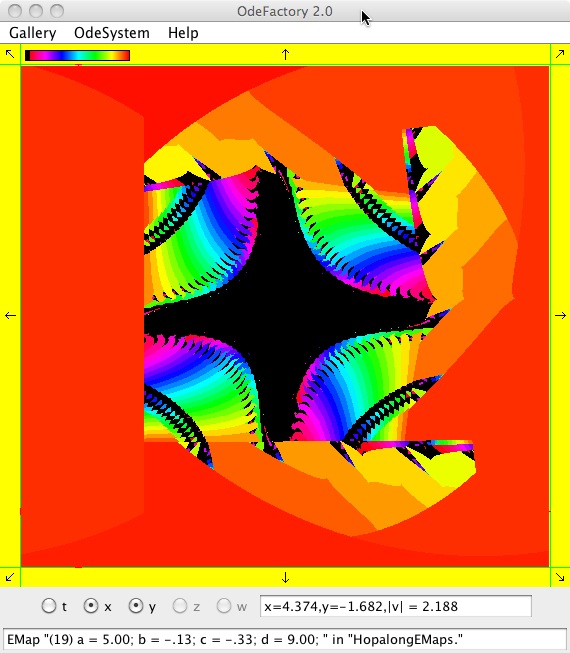
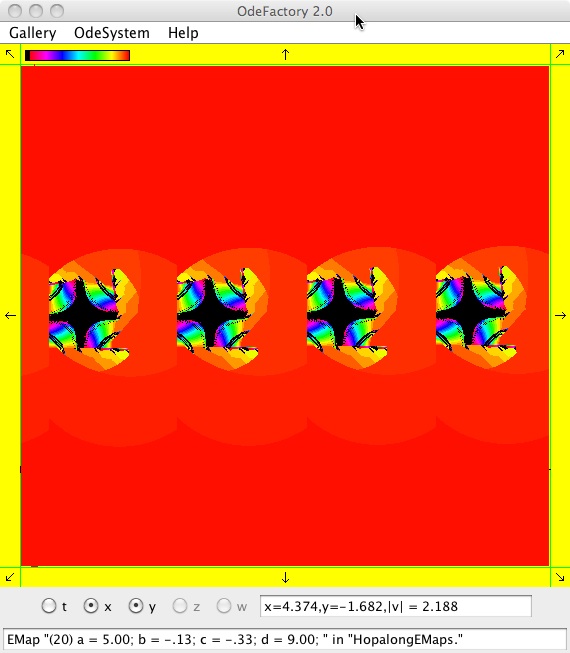
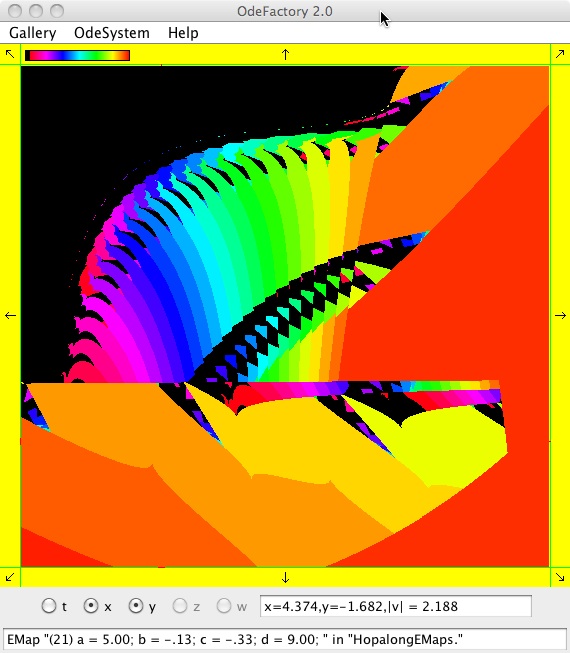


Here are some images that were added 3/6/13. The three 16-panel images were created by doing a right click in the EMap view to produce a shifted-pattern quilt image.